Graphics with Python: Difference between revisions
| (37 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
The Python matplotlib module | The Python matplotlib module adds tools for interactive 2-D and 3-D graphics to [http://prancer.physics.louisville.edu/astrowiki/index.php/Python_for_Physics_and_Astronomy our very short course in Python for scientific research]. Matplotlib provides for easily generating and saving plots in file formats you can display on the web or in other programs, print, and incorporate in documents. | ||
== Installation of matplotlib == | == Installation of matplotlib == | ||
| Line 25: | Line 25: | ||
The [http://matplotlib.org/users/index.html matplotlib on-line user's guide] offers a tutorial with many [http://matplotlib.org/examples/index.html examples], some of which we will look at here. The guide may also be [http://matplotlib | The [http://matplotlib.org/users/index.html matplotlib on-line user's guide] offers a tutorial with many [http://matplotlib.org/examples/index.html examples], some of which we will look at here. The guide may | ||
also be [http://prancer.physics.louisville.edu/classes/650/python/matplotlib/matplotlib-ref.pdf downloaded as a handy readable pdf] for off-line reference. There is also a helpful but unfinished [http://bespokeblog.wordpress.com/2011/07/05/basic-data-plotting-with-matplotlib-part-1-introduction/ quick start guide] written by an astrophysics graduate student. | |||
Let's look at a simple program that generates its own data and creates | === Creating one plot === | ||
Let's look at a simple program that generates its own data and creates one plot you can view on the screen using pyplot, a MATLAB-like interface: | |||
# Import the plotting and math packages | # Import the plotting and math packages | ||
| Line 49: | Line 52: | ||
amplitude.append(a) | amplitude.append(a) | ||
# Create an x-y plot of the data with labeled axes | # Create an x-y plot of the data with labeled axes | ||
plt.figure().canvas.set_window_title('Oscillator') | |||
plt.plot(time, amplitude) | plt.plot(time, amplitude) | ||
plt.xlabel('Time') | plt.xlabel('Time') | ||
| Line 59: | Line 63: | ||
Most of this program is used to create and prepare the data lists. The plotting is done in one line! We add labels to axes, a title to the plot, and show the work. The way in which it appears will depend on our installation, but the default is a Tkl interface that offers control for panning, zooming, and saving as png file. The data go into the plot as lists, and appear by default as a drawn line connecting the points. However, if you prefer red circles to a "pen down" line, then change the | Most of this program is used to create and prepare the data lists. The plotting is done in one line! We add labels to the canvas so the new window is labeled on the screen, to the axes, a title to the plot, and show the work. The way in which it appears will depend on our installation, but the default is a Tkl interface that offers control for panning, zooming, and saving as png file. The data go into the plot as lists, and appear by default as a drawn line connecting the points. However, if you prefer red circles to a "pen down" line, then change the plt.plot to | ||
plt.plot(time,amplitude,'ro') | |||
or to | or to | ||
| Line 114: | Line 118: | ||
# Show the data | # Show the data | ||
plt.show() | plt.show() | ||
This is what it looks like on the screen: | |||
<center> | |||
[[File:Damped_cos_screen.png]] | |||
</center> | |||
=== Saving the plot to a file === | |||
You may include a command to save a figure as a png file by adding | |||
fname = 'oscillator.png' | |||
plt.savefig(fname) | |||
The savefig function is sensitive to the file type in the extension, and | |||
fname = 'oscillator.ps' | |||
plt.savefig(fname,dpi=600) | |||
would be an example of a PostScript figure set to 600 dots per inch resolution. Supported formats are intended for high quality reproduction and include eps, ps, pdf, png, and svg, among others. The resolution may be controlled within the program, or in defaults for the user's custom startup file. | |||
=== Overplotting and separated plots === | |||
Matplotlib's ''pyplot.plot'' takes two arrays as arguments and is aware of several modifiers that determine how the data are plotted. When you use plot, you should be aware that implicitly it creates | |||
a figure and a subplot, and then uses these with the data you have provided. The subplot(nrows, ncols, plot_number) function allows you to make arrays of 9 or fewer plots on the same page. When you only have one plot, it is subplot(1,1,1). You are allowed to leave out the commas if the result is unambiguous, and you will sometimes see this written (lazily) as subplot(111). A second subplot on row 2 would be subplot(212) and so on. | |||
If the pyplot.plot function is repeated, it loads another set of x-y data into the one plot, each set with its own properties. The sets can be labeled, and the labels can be turned into legends in the plot. Here's a simple example: | |||
# Import the plotting and math packages | |||
import matplotlib.pyplot as plt | |||
import math | |||
# Define initial constants | |||
f0 = 5. | |||
tdecay = 2. | |||
# Create lists for the (x,y) data | |||
time = [] | |||
sine_amplitude = [] | |||
exp_amplitude = [] | |||
product_amplitude = [] | |||
# Calculate the data and append to the lists | |||
for i in range(0, 10000, 1): | |||
t = 0.001 * float(i) | |||
a1 = math.cos(2. * math.pi * f0 * t) | |||
a2 = math.exp(-t/tdecay) | |||
a = a1*a2 | |||
time.append(t) | |||
sine_amplitude.append(a1) | |||
exp_amplitude.append(a2) | |||
product_amplitude.append(a) | |||
# Create an x-y plot with labels | |||
plt.xlabel('Time') | |||
plt.ylabel('Amplitude') | |||
plt.title('A Damped Oscillator') | |||
plt.plot(time,exp_amplitude,'r.', label='Exponential') | |||
plt.plot(time, product_amplitude,'b-', label='Sine', linewidth=1.5) | |||
plt.legend() | |||
# Show the data | |||
plt.show() | |||
In this case we used abbreviations for the keywords, but it is clearer to use the full text and to write, for example | |||
plt.plot(time,product_amplitude, color='blue', linestyle = 'line', marker='None', | |||
label='Sine', linewidth=1.5) | |||
instead. | |||
You may create two separate plots instead of one, in the previous example, by using pyplot this way: | |||
# Create a figure | |||
fig = plt.figure() | |||
# Adjust white space between plots | |||
fig.subplots_adjust(hspace=0.5) | |||
# Create x-y plots of the data with labeled axes | |||
data1 = fig.add_subplot(2,1,1) | |||
plt.xlabel('Time') | |||
plt.ylabel('Amplitude') | |||
plt.title('Damping') | |||
data1.plot(time,exp_amplitude,'r.', label='Exponential') | |||
plt.legend() | |||
data2 = fig.add_subplot(2,1,2) | |||
plt.xlabel('Time') | |||
plt.ylabel('Amplitude') | |||
plt.title('A Damped Oscillator') | |||
data2.plot(time,product_amplitude, color='blue', linestyle='solid', marker='None', label='Sine', | |||
linewidth=1.5) | |||
plt.legend() | |||
# Show the data | |||
plt.show() | |||
We added subplots with the fig.add_subplot(a,b,c) function, which can also be written without the commas if there is no ambiguity in the numbers. The (a,b,c) mean an axb grid, and the cth subplot of that grid. So 111 is simply 1 plot, but 211 is a 2x1 grid and the first plot in the grid, while 212 is a 2x1 grid and the second plot. | |||
=== Other useful x-y data plotting functions === | |||
Other one-line plotting options that would create subplot(1,1,1) by default include | |||
import pyplot as plt | |||
plt.scatter | |||
plt.semilogx | |||
plt.semilogy | |||
plt.loglog | |||
Errorbars may be included with x and y data using | |||
plt.errorbar(x, y, yerr=None, xerr=None) | |||
It takes the x and y arrays, an array of y errors that defaults to ''None'', and an array of x errors that also defaults to ''None''. | |||
For more information on plot and other built-in plotting types, see the [http://matplotlib.org/api/pyplot_api.html?highlight=subplot#module-matplotlib.pyplot pyplot documentation]. | |||
=== Tick marks and data ranges === | |||
You've noticed that when data are loaded as a list or tuple, the plot function automatically selects the data range for both axes, and supplies tick marks. You may add minor ticks and control their properties | |||
with | |||
plt.minorticks_on() | |||
plt.tick_params(which='major', length=7) | |||
plt.tick_params(which='minor', length=4, color='r') | |||
where the first one turns on the minor ticks (minorticks_off() is the default), and the others change their length and color. | |||
You may also limit the range of data shown in the plot and retain the automatic tick marking by adding functions such as these ''after creating the plot'' for the example show above | |||
oscillator = plt.plot(time,amplitude) | |||
plt.xlim(-10.,10.) | |||
plt.ylim(-200.,200.) | |||
With these limits, the range of x shown will be from -10 to +10 instead of the range in the x-data, and y will be shown from -200 to +200. | |||
== Interactive plotting == | |||
It is often useful to view a function or processing operation interactively with a graph that shows the results of changing one or more variables. This is an example written by Michael Tague, a student | |||
in the Research Methods class in 2013. We begin by setting up matplotlib and numpy in the usual way: | |||
import matplotlib.pyplot as plt | |||
import numpy as np | |||
from matplotlib.widgets import Slider | |||
and you see in the last line we have added a useful "Slider" control. Next we create a series of time values as a numpy array | |||
t = np.arange(0, 1, 0.001) | |||
from 0 to 1 in steps of 0.001 which we take to be seconds. From this we create a matching series of amplitudes | |||
y = np.sin(2*np.pi*t*f)/np.exp(t*d) | |||
and a plot | |||
p, = plt.plot(t,y) | |||
You could stop here with "plt.show()" and be done with simple plot of the sinusoidal function. If you want to add controls to make the plot interactive, then rather than showing the plot now | |||
add adjustable subplots. The command | |||
plt.subplots_adjust(bottom=0.25) | |||
makes room for the sliders at the bottom of the plot. We add axes | |||
fax = plt.axes([0.25, 0.14, 0.5, 0.03]) | |||
and a frequency Slider | |||
fs = Slider(fax, 'Freq', 0.0, 10.0, valinit=f) | |||
Now we create a function that updates the frequencies whenever the slider is changed | |||
def f_update(val): | |||
global f, p | |||
f = val | |||
y = np.sin(2*np.pi*t*f)/np.exp(t*d) | |||
p.set_ydata(y) | |||
plt.draw(); | |||
fs.on_changed(f_update) | |||
The decay constant slider is the same as this, but for the decay, d, we move it down a bit. | |||
dax = plt.axes([0.25, 0.07, 0.5, 0.03]) | |||
ds = Slider(dax, 'Decay', 0.0, 10.0, valinit=d) | |||
def d_update(val): | |||
global d | |||
d = val | |||
y = np.sin(2*np.pi*t*f)/np.exp(t*d) | |||
p.set_ydata(y) | |||
plt.draw(); | |||
ds.on_changed(d_update) | |||
Now we show the interactive plot with | |||
plt.show() | |||
[[File:Interactive_damped_oscillator.png | center | 600px]] | |||
== A little 3D plotting == | |||
One of the features that makes Mathematica and Matlab so useful is their built-in 3D plotting for multivariable surfaces and objects. Matplotlib does this very easily in Python, but the features are so rich we can only sample a few of them here. | |||
As an appetizer, consider surface3d_demo2.py from matplotlib: | |||
from mpl_toolkits.mplot3d import Axes3D | |||
import matplotlib.pyplot as plt | |||
import numpy as np | |||
fig = plt.figure() | |||
ax = fig.add_subplot(111, projection='3d') | |||
u = np.linspace(0, 2 * np.pi, 100) | |||
v = np.linspace(0, np.pi, 100) | |||
x = 10 * np.outer(np.cos(u), np.sin(v)) | |||
y = 10 * np.outer(np.sin(u), np.sin(v)) | |||
z = 10 * np.outer(np.ones(np.size(u)), np.cos(v)) | |||
ax.plot_surface(x, y, z, rstride=4, cstride=4, color='b') | |||
plt.show() | |||
The first line imports a toolkit that provides 3d projection. For this one, you would need the mpl_tookits package. | |||
The second line imports matplotlib.pyplot and uses plt to stand for it in our code. | |||
The third line imports numpy, numerical Python, which we are going to look at in more detail later. Here, it provides a way to handle the data transparently even if you do not know how the code works yet. | |||
We create a figure called "fig", and we add one and only one subplot "ax" that will hold our work. Notice that the subplot is a "3d" projection. | |||
Numpy is used to create two linear arrays, one from 0 to 2 Pi that we think of as Phi, the azimuth angle in spherical coordinates. The other, from 0 to Pi, is Theta, the altitude of spherical coordinates. From these we calculate a set of x, y, and z that are the coordinates of points on a surface. Here, "np.outer" means "outer product" of two linear arrays, that is each of x, y, and z are matrices that contain the coordinates (x,y,z) of points on the unit sphere or radius 10 selected by the angles u and v. | |||
The plotting is done in one line from the mpl toolkit with plot_surface(), and then dislayed for us to use with show(). | |||
From that little program we get an interactive 3D display of a sphere that looks like this: | |||
[[File:Matplotlib_3d_sphere.png | center | 600px]] | |||
There's a [http://matplotlib.org/mpl_toolkits/mplot3d/tutorial.html 3D tutorial for matplotlib from their website] illustrating other examples. | |||
== Examples == | |||
For examples of Python illustrating plotting with matplotlib, see the [http://prancer.physics.louisville.edu/astrowiki/index.php/Python_examples examples] section. | |||
== Assignments == | |||
For the assigned homework to use these ideas, see the [http://prancer.physics.louisville.edu/astrowiki/index.php/Python_assignments assignments] section. | |||
Revision as of 21:03, 27 January 2015
The Python matplotlib module adds tools for interactive 2-D and 3-D graphics to our very short course in Python for scientific research. Matplotlib provides for easily generating and saving plots in file formats you can display on the web or in other programs, print, and incorporate in documents.
Installation of matplotlib
The current version 1.2 may be included in some Linux distributions. Version 1.1 has most of the features you will need now, and it is in Ubuntu and OpenSuse packages that can be added to your core Python system after you also install numpy. For example, under Ubuntu you would use
sudo apt-get install python-matplotlib
to get the most recent version available for your system and resolve missing components.
For Windows and MacOS users, if you installed the Enthought version of Python you will have it "out of the box". For others, look at the matplotlib installation website for directions on how to install it. You will need numpy too, and it also comes in the Enthought collection.
Once you have it installed, programs that use this library will have to import it with lines such as
import numpy as np import matplotlib as plt
to make the functions available. With these, numpy functions will start with np. and mathplotlib functions will have plt. in front of the function name, which shortens the code you would write. You can check that your computer has numpy and matplotlib by trying these commands in interactive Python or Idle. The version numbers will be available too with
print np.__version__ print plt.__version__
Learning the basics of 2D data and function plotting
The matplotlib on-line user's guide offers a tutorial with many examples, some of which we will look at here. The guide may also be downloaded as a handy readable pdf for off-line reference. There is also a helpful but unfinished quick start guide written by an astrophysics graduate student.
Creating one plot
Let's look at a simple program that generates its own data and creates one plot you can view on the screen using pyplot, a MATLAB-like interface:
# Import the plotting and math packages import matplotlib.pyplot as plt import math
# Define initial constants f0 = 5. a0 = 100. tdecay = 2.
# Create lists for the (x,y) data time = [] amplitude = []
# Calculate the data and append to the lists for i in range(0, 10000, 1): t = 0.001 * float(i) a = a0 * math.exp(-t/tdecay)*math.cos(2. * math.pi * f0 * t) time.append(t) amplitude.append(a)
# Create an x-y plot of the data with labeled axes
plt.figure().canvas.set_window_title('Oscillator')
plt.plot(time, amplitude)
plt.xlabel('Time')
plt.ylabel('Amplitude')
plt.title('A Damped Oscillator')
# Show the data
plt.show()
Most of this program is used to create and prepare the data lists. The plotting is done in one line! We add labels to the canvas so the new window is labeled on the screen, to the axes, a title to the plot, and show the work. The way in which it appears will depend on our installation, but the default is a Tkl interface that offers control for panning, zooming, and saving as png file. The data go into the plot as lists, and appear by default as a drawn line connecting the points. However, if you prefer red circles to a "pen down" line, then change the plt.plot to
plt.plot(time,amplitude,'ro')
or to
'r--' # red dashes 'bs' # blue stars 'g^' # green triangles
The properties of the line would be controlled by variables in the plot function using MATLAB-style string/value pairs.
plt.plot(time,amplitude, color='g', linewidth='2.0')
or alternatively a plot control function
lines = plt.plot(time,amplitude) plt.setp(lines, color='r', linewidth=2.0)
This has the useful feature that interactively plt.setp(lines) will show you all the parameters and their values.
The data in this example are input as lists, but they could be tuples instead:
# Import the plotting and math packages import matplotlib.pyplot as plt import math
# Define initial constants f0 = 5. a0 = 100. tdecay = 2.
# Create lists for the (x,y) data time = [] amplitude = []
# Calculate the data and append to the lists for i in range(0, 10000, 1): t = 0.001 * float(i) a = a0 * math.exp(-t/tdecay)*math.cos(2. * math.pi * f0 * t) time.append(t) amplitude.append(a)
x = tuple(time) y = tuple(amplitude)
# Create an x-y plot of the data with labeled axes
plt.xlabel('Time')
plt.ylabel('Amplitude')
plt.title('A Damped Oscillator')
oscillator = plt.plot(time,amplitude)
plt.setp(oscillator, color='m', linewidth=1.5)
# Show the data
plt.show()
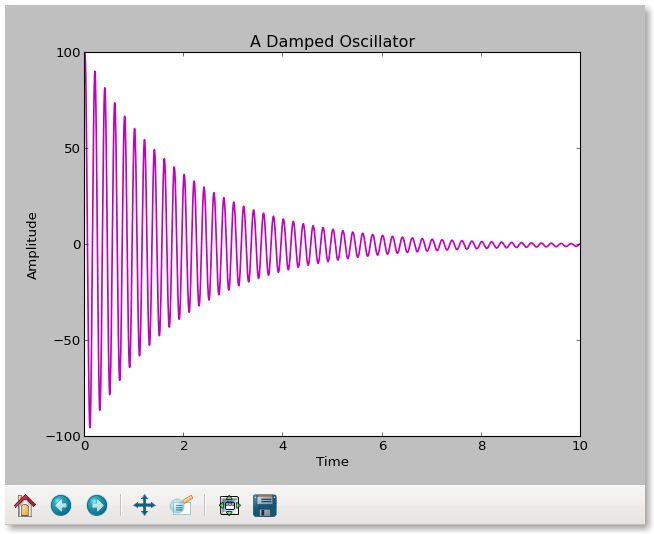
This is what it looks like on the screen:
Saving the plot to a file
You may include a command to save a figure as a png file by adding
fname = 'oscillator.png' plt.savefig(fname)
The savefig function is sensitive to the file type in the extension, and
fname = 'oscillator.ps' plt.savefig(fname,dpi=600)
would be an example of a PostScript figure set to 600 dots per inch resolution. Supported formats are intended for high quality reproduction and include eps, ps, pdf, png, and svg, among others. The resolution may be controlled within the program, or in defaults for the user's custom startup file.
Overplotting and separated plots
Matplotlib's pyplot.plot takes two arrays as arguments and is aware of several modifiers that determine how the data are plotted. When you use plot, you should be aware that implicitly it creates a figure and a subplot, and then uses these with the data you have provided. The subplot(nrows, ncols, plot_number) function allows you to make arrays of 9 or fewer plots on the same page. When you only have one plot, it is subplot(1,1,1). You are allowed to leave out the commas if the result is unambiguous, and you will sometimes see this written (lazily) as subplot(111). A second subplot on row 2 would be subplot(212) and so on.
If the pyplot.plot function is repeated, it loads another set of x-y data into the one plot, each set with its own properties. The sets can be labeled, and the labels can be turned into legends in the plot. Here's a simple example:
# Import the plotting and math packages import matplotlib.pyplot as plt import math
# Define initial constants f0 = 5. tdecay = 2.
# Create lists for the (x,y) data time = [] sine_amplitude = [] exp_amplitude = [] product_amplitude = []
# Calculate the data and append to the lists for i in range(0, 10000, 1): t = 0.001 * float(i) a1 = math.cos(2. * math.pi * f0 * t) a2 = math.exp(-t/tdecay) a = a1*a2 time.append(t) sine_amplitude.append(a1) exp_amplitude.append(a2) product_amplitude.append(a)
# Create an x-y plot with labels
plt.xlabel('Time')
plt.ylabel('Amplitude')
plt.title('A Damped Oscillator')
plt.plot(time,exp_amplitude,'r.', label='Exponential')
plt.plot(time, product_amplitude,'b-', label='Sine', linewidth=1.5)
plt.legend()
# Show the data plt.show()
In this case we used abbreviations for the keywords, but it is clearer to use the full text and to write, for example
plt.plot(time,product_amplitude, color='blue', linestyle = 'line', marker='None', label='Sine', linewidth=1.5)
instead.
You may create two separate plots instead of one, in the previous example, by using pyplot this way:
# Create a figure fig = plt.figure()
# Adjust white space between plots fig.subplots_adjust(hspace=0.5)
# Create x-y plots of the data with labeled axes
data1 = fig.add_subplot(2,1,1)
plt.xlabel('Time')
plt.ylabel('Amplitude')
plt.title('Damping')
data1.plot(time,exp_amplitude,'r.', label='Exponential')
plt.legend()
data2 = fig.add_subplot(2,1,2)
plt.xlabel('Time')
plt.ylabel('Amplitude')
plt.title('A Damped Oscillator')
data2.plot(time,product_amplitude, color='blue', linestyle='solid', marker='None', label='Sine',
linewidth=1.5)
plt.legend()
# Show the data plt.show()
We added subplots with the fig.add_subplot(a,b,c) function, which can also be written without the commas if there is no ambiguity in the numbers. The (a,b,c) mean an axb grid, and the cth subplot of that grid. So 111 is simply 1 plot, but 211 is a 2x1 grid and the first plot in the grid, while 212 is a 2x1 grid and the second plot.
Other useful x-y data plotting functions
Other one-line plotting options that would create subplot(1,1,1) by default include
import pyplot as plt plt.scatter plt.semilogx plt.semilogy plt.loglog
Errorbars may be included with x and y data using
plt.errorbar(x, y, yerr=None, xerr=None)
It takes the x and y arrays, an array of y errors that defaults to None, and an array of x errors that also defaults to None.
For more information on plot and other built-in plotting types, see the pyplot documentation.
Tick marks and data ranges
You've noticed that when data are loaded as a list or tuple, the plot function automatically selects the data range for both axes, and supplies tick marks. You may add minor ticks and control their properties with
plt.minorticks_on() plt.tick_params(which='major', length=7) plt.tick_params(which='minor', length=4, color='r')
where the first one turns on the minor ticks (minorticks_off() is the default), and the others change their length and color.
You may also limit the range of data shown in the plot and retain the automatic tick marking by adding functions such as these after creating the plot for the example show above
oscillator = plt.plot(time,amplitude) plt.xlim(-10.,10.) plt.ylim(-200.,200.)
With these limits, the range of x shown will be from -10 to +10 instead of the range in the x-data, and y will be shown from -200 to +200.
Interactive plotting
It is often useful to view a function or processing operation interactively with a graph that shows the results of changing one or more variables. This is an example written by Michael Tague, a student in the Research Methods class in 2013. We begin by setting up matplotlib and numpy in the usual way:
import matplotlib.pyplot as plt import numpy as np from matplotlib.widgets import Slider
and you see in the last line we have added a useful "Slider" control. Next we create a series of time values as a numpy array
t = np.arange(0, 1, 0.001)
from 0 to 1 in steps of 0.001 which we take to be seconds. From this we create a matching series of amplitudes
y = np.sin(2*np.pi*t*f)/np.exp(t*d)
and a plot
p, = plt.plot(t,y)
You could stop here with "plt.show()" and be done with simple plot of the sinusoidal function. If you want to add controls to make the plot interactive, then rather than showing the plot now add adjustable subplots. The command
plt.subplots_adjust(bottom=0.25)
makes room for the sliders at the bottom of the plot. We add axes
fax = plt.axes([0.25, 0.14, 0.5, 0.03])
and a frequency Slider
fs = Slider(fax, 'Freq', 0.0, 10.0, valinit=f)
Now we create a function that updates the frequencies whenever the slider is changed
def f_update(val): global f, p f = val y = np.sin(2*np.pi*t*f)/np.exp(t*d) p.set_ydata(y) plt.draw();
fs.on_changed(f_update)
The decay constant slider is the same as this, but for the decay, d, we move it down a bit.
dax = plt.axes([0.25, 0.07, 0.5, 0.03]) ds = Slider(dax, 'Decay', 0.0, 10.0, valinit=d)
def d_update(val): global d d = val y = np.sin(2*np.pi*t*f)/np.exp(t*d) p.set_ydata(y) plt.draw();
ds.on_changed(d_update)
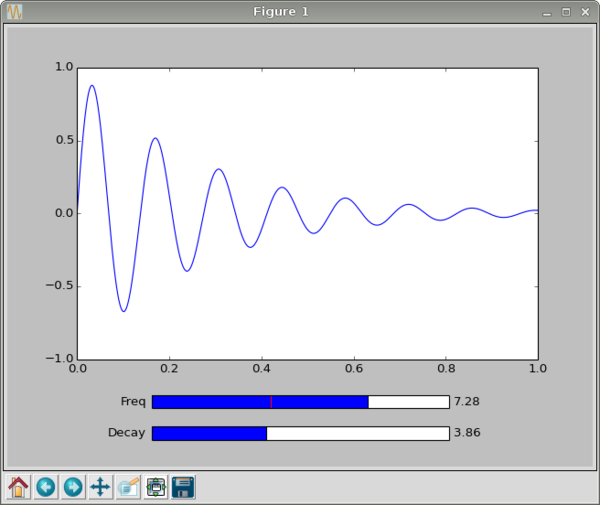
Now we show the interactive plot with
plt.show()
A little 3D plotting
One of the features that makes Mathematica and Matlab so useful is their built-in 3D plotting for multivariable surfaces and objects. Matplotlib does this very easily in Python, but the features are so rich we can only sample a few of them here.
As an appetizer, consider surface3d_demo2.py from matplotlib:
from mpl_toolkits.mplot3d import Axes3D import matplotlib.pyplot as plt import numpy as np
fig = plt.figure() ax = fig.add_subplot(111, projection='3d')
u = np.linspace(0, 2 * np.pi, 100) v = np.linspace(0, np.pi, 100)
x = 10 * np.outer(np.cos(u), np.sin(v)) y = 10 * np.outer(np.sin(u), np.sin(v)) z = 10 * np.outer(np.ones(np.size(u)), np.cos(v)) ax.plot_surface(x, y, z, rstride=4, cstride=4, color='b')
plt.show()
The first line imports a toolkit that provides 3d projection. For this one, you would need the mpl_tookits package.
The second line imports matplotlib.pyplot and uses plt to stand for it in our code.
The third line imports numpy, numerical Python, which we are going to look at in more detail later. Here, it provides a way to handle the data transparently even if you do not know how the code works yet.
We create a figure called "fig", and we add one and only one subplot "ax" that will hold our work. Notice that the subplot is a "3d" projection.
Numpy is used to create two linear arrays, one from 0 to 2 Pi that we think of as Phi, the azimuth angle in spherical coordinates. The other, from 0 to Pi, is Theta, the altitude of spherical coordinates. From these we calculate a set of x, y, and z that are the coordinates of points on a surface. Here, "np.outer" means "outer product" of two linear arrays, that is each of x, y, and z are matrices that contain the coordinates (x,y,z) of points on the unit sphere or radius 10 selected by the angles u and v.
The plotting is done in one line from the mpl toolkit with plot_surface(), and then dislayed for us to use with show().
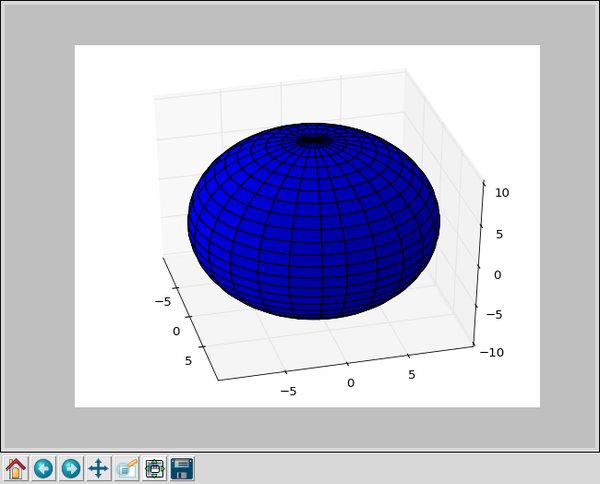
From that little program we get an interactive 3D display of a sphere that looks like this:
There's a 3D tutorial for matplotlib from their website illustrating other examples.
Examples
For examples of Python illustrating plotting with matplotlib, see the examples section.
Assignments
For the assigned homework to use these ideas, see the assignments section.